| Up Contents | ||||||||
Force Field development1,2-DifluoroethaneAs an example, let us construct a force field compatible with the AMBER94 for 1,2-difluoroethane. Draw the molecule with a mouse
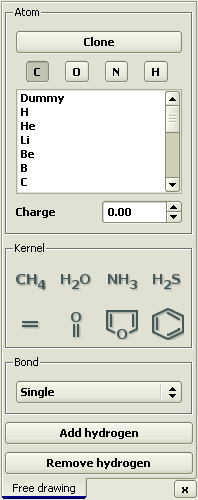
Open the Free drawing panel in the Build > Free drawing
menu or by pressing the toolbar button
Select methane as a primer for the model by clicking the
Right-click the hydrogen atom to transform it to carbon.
Click Add hydrogen button.
Choose fluoride from the list of elements and transform two hydrogen atoms into it by right-clicking them.
A crude model of 1,2-difluoroethane is ready.
Let us transform it into two models in gauche- and trans-conformations.
To construct a gauche model, invoke the Geometry Editor.
Conformation refinement
The resulting model has a gauche conformation but wrong bond lengths.
Rapid preliminary optimization can be done by semiempirical
quantum chemical methods. Open the GAMESS panel in the
Tools > PC GAMESS or by pressing the toolbar button
The conformation can be further specified using Ab inition calculations. Let us first use the 3-21+G basis with an electrons correlation at the MP2 level.
Then, perform the optimization in MP2 / 6-311+G(2d,p). AMBER94 was developed in the basis of 6-31G(d), but we will shift to MP2 / 6-311+G(2d,p) since the 2d bases provide for substantially better results, and current computers are powerful enough to make such calculations routine. An MP2 / 6-311+G(2d,p) optimization yields an accurate model in a gauche conformation. Save it to a file.
Potential-derived charges on atomsWith a model in a sensible conformation, one can determine the partial charges on atoms. Since our molecule includes only four heavy atoms, the electrostatic field around it can be calculated using a relatively large basis, e.g., aug-cc-pVTZ. For typical calculations, aug-cc-pVDZ, can be recommended since it yields the dipole moments close to experimental (for MP2 electron correlation). Select the Electrostatics calculation, MP2 correlation, and aug-cc-pVTZ basis.
After a calculation for about 1 hour, partial charges are assigned to the model atoms. Do not forget to save the model after the calculation. C 0.0880 C 0.0873 H 0.0908 H 0.0560 H 0.0910 H 0.0561 F -0.2347 F -0.2345 Trans-1,2-difluoroethaneRepeat the above steps to generate a model of difluoroethane in a trans conformation. C 0.1236 C 0.1236 H 0.0730 H 0.0728 H 0.0728 H 0.0730 F -0.2694 F -0.2694 The charges on atoms in the gauche and trans models slightly differ. Fortunately, the differences in this case are not great and we can just average the charges on equivalent atoms in both conformations. C 0.1056 C 0.1056 H 0.0732 H 0.0732 H 0.0732 H 0.0732 F -0.2520 F -0.2520 The conformational dependence of charges is a serious problem of parametrization of molecular mechanics force fields. Some kind of averaging is a common solution. Parameters of valence interactionsConstruction of an accurate model includes a good torsion energy approximation. This requires either valid experimental data or adequate quantum chemical calculations. The torsion potentials cannot be specified by analogy with other molecules, such analogies are not practicable. The torsion potentials should be specified obligatorily after the partial charges on atoms are specified, since they heavily depend the charges. Let us calculate our optimized gauche and trans models in MP4 / aug-cc-pVDZ. Select the aug-cc-pVDZ bases in the Standard section, specify MP4 in the above panel, and select the ENERGY calculation. Do not forget to test the settings by the Check button, and start calculation. (The calculation lasted 13 min on our PC).
Thus, at the MP4 / aug-cc-pVDZ approximation, the gauche conformation proved more stable than the trans conformation by -0.0010739464 Hartree (-0.674 kcal/mol or -2.82 kJ/mol). The experimental values are 0.83 kcal/mol (S. Nonoyama et al., 51st Annual Meeting of Chemical Society, Kanazawa, Japan) and 0.57 ± 0.09 kcal/mol, (K.B. Wiberg and M.A. Murcko, J. Phys. Chem., 1987, vol. 91). Before selecting the force field parameters, we have to specify molecular mechanics types for the atoms. Force fields by different authors have different sets of types. Fine Types were introduced to allow switching between them. Fine Types are the most specialized types for all supported force fields. The /data/ForceField folder contains the FineType.mlm file with a table of conversion from Fine Types to particular force field types. Unique types can be created for all atoms in the molecule, but we will use the -CH2- and H2C- types for carbon and hydrogen, respectively. Create a new Fine Type F-tutor for fluorine. We specified that it should be converted to the F type of the AMBER94 field. Restart the program to apply the FineType.mlm file changes. Save the corresponding types to the model files. The result can look as follows: Let us open the model and calculate its energy. A warning appears "No such angle parameter: CT CT F", pointing to missing parameters for the current force field (AMBER94). The full list of missing parameters is specified in /Report/FF_error.txt. Another way to determine the missing parameters is to save the model in the mmol format. Specify the missing parameters in the /data/ForceField/AMBER94_2_angle.xls file. In this example, they will be chosen by analogy with those available in AMBER94. For an accurate model, they can be determined by the approximation of quantum mechanical calculations; however, they are not critical for many tasks. Specify the following parameters: Energy calculation finds no unspecified parameters. Although the torsion angles have not been specified, AMBER94 has default values for the *-CT-CT-* angles. We can specify the model by setting the F-CT-CT-F value. Include the following line in the /data/ForceField/AMBER94_3_torsional.xls file: This line should follow the lines with asterisks to override the default value. The force constant was temporarily set to unity. Its value should be selected so that the difference between the trans and gauche conformation energies approximates that obtained by quantum mechanics or experiment. The comparison should be performed in local energy minima; accordingly, both models should be optimized after each parameter change. The Optimization panel is invoked from the Compute > Optimize menu. The default parameters are suitable for our task. Several experiments with the force constant demonstrate that the V/2 value of 2.02 kcal/mol provides for the desired energy difference. As a result, a simple model of 1,2-difluoroethane was constructed using the following steps:
A finer model requires the calibration of van der Waals and electrostatic interactions to fit the vaporization heat and density of liquid 1,2-difluoroethane to experimental data.
|
||||||||
| Up Contents | ||||||||
| Copyright © 2006-2008 Agile Molecule. All rights reserved | ||||||||